Junk DNA: A Journey Through the Dark Matter of the Genome (10 page)
Read Junk DNA: A Journey Through the Dark Matter of the Genome Online
Authors: Nessa Carey

This particular stretch of junk DNA is called the centromere, and unlike the telomeres from the last chapter, the centromere is found on the interior of a chromosome. Depending on the chromosome, it may be pretty much in the middle, or it may be near to an end. Its position is consistent in the sense that on human chromosome 1, for example, it’s always near the middle whereas in human chromosome 14 it’s always near the end.
Centromeres are essentially attachment points for a set of proteins that drag the separated chromosomes to opposite ends of the cell. Imagine Spider-Man is standing in a set position and needs to get something. He throws a web at the thing he wants, and then drags it to him. Now imagine that a very tiny Spider-Man is standing at one end of a cell. He throws a web at the chromosome he wants, the web attaches, and he pulls the chromosome to
his end of the cell. A tiny Spider-Man clone does the same thing at the opposite end of the cell for the other chromosome in the matching pair.

Figure 6.2
This shows the cell division process that generates gametes (eggs or sperm) each containing just one of every pair of chromosomes. The process initially looks like the standard cell division shown in Figure 6.1. However, this is followed by a second separation of chromosome pairs, to create gametes with only half the normal number of chromosomes. There is also an early event where genetic material is swapped over within chromosome pairs, to create greater genetic diversity in offspring, but this isn’t shown in this figure.
There is a complication for Spider-Man. Most of the surface of the chromosome is coated with web-repellent. There is only one part where his web will stick. This part is the centromere. In the cell the centromere attaches to a long string of proteins which pulls the chromosome away from the centre and to the periphery. This string of proteins is called the spindle apparatus.
Centromeres play a very important and consistent role in all species. They form the essential attachment point for the spindle apparatus. It’s essential that this system works properly, or cell division goes wrong. Given that this is such a vital process, we would expect that the centromere DNA sequence would be highly conserved throughout the evolutionary tree. But weirdly, this isn’t the case at all. Once we move beyond yeast
a
and microscopic worms,
b
the DNA sequence is highly variable when we look at different species.
2
In fact, the DNA sequence of a centromere may differ between two chromosomes in the same cell. This level of sequence diversity, in the face of functional consistency, is really quite counterintuitive. Happily, we are starting to understand how this vital region of junk DNA manages to pull off this strange evolutionary trick.
In human chromosomes, the centromeres are formed from repeats of a DNA sequence that is 171 base pairs in length.
c
These 171 base pairs are repeated over and over, and may reach lengths of up to 5 million bases in total.
3
The critical feature of the centromere is that it acts as a location for the binding of the protein called CENP-A (
Cen
tromeric
P
rotein-
A
).
4
The CENP-A gene is highly conserved between species, in contrast to the centromere DNA.
Our Spider-Man analogy might be useful again here in terms of understanding the apparent evolutionary conundrum we laid out earlier. Spidey’s web can bind to CENP-A protein. It doesn’t matter if the CENP-A protein is bound to meat, bricks, potatoes or lightbulbs. So long as the CENP-A protein is bound to something, Spider-Man’s web will stick to it, and pull the CENP-A and the something towards our superhero.
So, the DNA sequence at the centromere can vary enormously between species, ranging from meat to lightbulbs. What matters is that the CENP-A protein remains the same, so that the highly conserved spindle apparatus can stick to it and pull the chromosomes apart to opposite poles of the dividing cell.
CENP-A isn’t the only protein that is found at the centromere; many others are also present. It’s possible to knock out the expression of CENP-A in cells in the laboratory. When this happens, the other proteins that should bind to the centromere stop doing so.
5
,
6
However, when the experiment is performed the other way around – knocking out expression of one of the other proteins – CENP-A continues to bind at the centromere.
7
This demonstrates that CENP-A acts as a foundation stone.
When researchers over-expressed CENP-A in cells from fruit flies, they found that the chromosomes began to create centromeres in unusual positions.
8
But the situation in human cells seems to be more complicated, because over-expression of CENP-A doesn’t result in new, abnormally located centromeres.
9
It seems that in humans, CENP-A is necessary for centromere formation, but it’s not sufficient.
The CENP-A acts as the essential cornerstone for the recruitment of all the other proteins that are also required for the spindle apparatus to do its job. When a cell is actively dividing, over 40 different proteins build up from the CENP-A. They do so in a step-wise fashion, like adding on LEGO bricks in a particular order. Immediately after the duplicated chromosomes have been pulled to the opposite ends of the cell, this big complex falls apart again. This whole process can take less than an hour. We don’t know what controls all of this, but some of it is down to a simple physical feature. Normally, the nucleus has a membrane around it, and large protein molecules find it really difficult to get through this. When the cell is ready to separate its replicated chromosomes, this barrier breaks down temporarily and the proteins can join on
to the complex at the centromere.
10
It’s like having a removal company outside your house. They are ready to shift your furniture but can’t get on with the job unless you open the door and let them in.
Location, location, location
We are still left with a difficult conceptual problem. If the DNA sequence at the centromere isn’t very conserved, and the critical factor is the placement of the CENP-A protein, how does the cell ‘know’ where the centromere should be on each chromosome? Why is it always near the middle of chromosome 1, but near the end of chromosome 14?
To understand this, we have to develop a more sophisticated image of the DNA in our cells. The DNA double helix is an iconic image, probably the defining image in biology. But it doesn’t really represent what DNA is like. DNA is a very long spindly molecule. If you stretched out the DNA from one human cell it would reach for two metres, assuming you joined up the material from all the chromosomes. But this DNA has to fit into the nucleus of a cell, and the nucleus has a diameter of just one hundredth of a millimetre.
This is like trying to fit something that is the vertical height of Mount Everest into a capsule the size of a golf ball. If you are trying to fit a climbing rope the height of Mount Everest into a golf ball, that clearly won’t work. On the other hand, if you replace the climbing rope with a filament thinner than a human hair, you’ll probably be OK.
Although human DNA is long, it’s very thin, so it is possible to fit it into the nucleus. But there is, as always, a complication. It’s not enough just to jam the DNA into a small space. The easiest way to visualise why not is to think of strings of Christmas tree lights. If at the end of the festive season you take the lights off the tree and shove them into a box, they will take up a lot of space.
You will also almost certainly find that when you come to use them again the following year they are all tangled up. It will take you ages to unravel them and there is a fair chance you will break some of them. In their tangled state, you would also really struggle to get to just one particular bulb.
But, if you are a freakishly organised person, you will wrap each string of lights around a piece of cardboard before storing them away. And your organisational acumen will be rewarded next Christmas when you take the lights out of the surprisingly small box you were able to use for storage. Not only did you save on loft space, you also will find that it’s very easy to unwind the lights, none of the strands get tangled around each other or snapped, and you can access your one favourite bulb very easily.
The same process happens in our cells. DNA is not stored as a random bundle of scrunched-up genetic material. Instead it is wrapped around certain proteins. This stops the DNA getting tangled and broken, allows it to be squeezed in an orderly fashion into a small space, and also keeps it structured so that the cell can access different regions as necessary, in order to switch individual genes on or off.
The DNA in our cells is wrapped around particular proteins, called histones. The basic structure is shown in Figure 6.3. Eight histone proteins – two each of four different types – form an octamer. DNA wraps around this octamer, like a skipping rope around eight tennis balls. There are huge numbers of these octamers all along our genome.
CENP-A is a close cousin of one of these histone proteins, sharing much of the same amino acid sequence, but with some important differences. At the centromere, both copies of one of the standard histone proteins are missing,
d
and CENP-A is present in the octamer instead, as shown in Figure 6.4.
11
There are thousands
of these octamers containing CENP-A at the centromere of each chromosome.

Figure 6.3
DNA, represented by the solid black band, is wrapped around packages of eight histone proteins (two each of four different types).
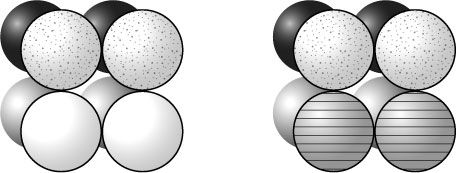
Figure 6.4
The octamer of histone proteins on the left represents the standard arrangement found throughout most of the genome. The octamer on the right represents the specialised octamers found at the centromeres. One of the standard pairs of histone proteins has been replaced by a pair of specialised centromere histone proteins, called CENP-A. These are represented by the striped globes.
The CENP-A in these thousands of octamers at the centromeres gives the spindle apparatus something to hang on to, when it’s trying to pull the chromosomes apart. One of the effects of inserting CENP-A into the octamers is that it makes the centromere regions more rigid.
12
If we think about trying to pull a
blob of jelly, compared with a boiled sweet, it’s obvious that the increased rigidity will be an advantage for the actions of the spindle apparatus.
But we still keep coming back to the same problem. Why is CENP-A inserted into the octamers at the centromere, but not at other regions? This isn’t driven by the DNA sequence. Other regions of our genome also contain junk DNA with similar sequences to those found at the centromeres, but CENP-A doesn’t accumulate at these.
13
CENP-A is only found at centromeres, but in some ways it’s the presence of CENP-A that actually defines what a centromere is. How have human cells evolved in such a way that an inherently unstable situation has led to complete genetic stability in terms of cell division?
